What is the
significance of the results of this research?
All the cells in our organism have the same
genome codified in the DNA. But we all know that not all the cells of the
organism are the same: a nervous cell, for example, is very different from a
blood cell or a muscle cell. The difference is due to the fact that each type
of cell has the capability to activate a different genetic programme, turning
on or off particular genes in various stages of embryonic development and of
adult-life. Researchers describe this phenomenon with a very appropriate term:
in fact, they say that each cell expresses
different genes. This research has identified, for the first time, the expression-profile of human chromosome
21 genes, that is to say, a catalogue of all the genes of chromosome 21, which
are either on or off in different tissues and at various stages of embryonic
development. The outcome is the equivalent of an anatomic atlas, so it is
possible to understand where (= in which tissue) and when (= at what stage of
development) the chromosome 21 genes are expressed.
Why do we need an
atlas of gene expression?
The knowledge that a gene is “on”, or
expressed, at a particular stage of embryonic development, in a specific tissue
or organ, may indicate that this gene has an important role
in developmental regulation or a function in that tissue or organ. The same is
true for pathological situations: if a gene is differentially expressed in a
pathological cell compared to a healthy cell, this suggests that this same gene
might be implicated in the origins of the disease.
This study provides significant information on
which genes of chromosome 21 could be important for a correct development of
various organs.
Why has Chromosome 21,
in particularly, been studied?
Humans have 23 pairs of chromosomes, generally
numbered from the largest to the smallest. Thus, chromosome 21 is one of the
smallest chromosomes - containing approximately 200 genes - but its study is of
enormous importance for the research on Down Syndrome
and in a considerable number of other genetic diseases. In fact, an extra third
copy of chromosome 21 is present in humans affected by Down Syndrome
(instead of two copies, which is normally the case). Of the approximately 200
genes of Chromosome 21 at least 30 are involved in as many genetic diseases,
such as, myopathy, anemia, blood platelet disorders, sclerosis, heart rhythm
problems, one form of epilepsy, at least two forms of deafness and two of
cataracts. Furthermore, the APP gene is localized on chromosome 21, and this is
one gene involved in Alzheimer disease.
What is the difference between the expression atlas and the map of the Human Genome published last year?
The Genome map is the sequence of all the “letters” of the DNA, composing the genome. Here, researchers can find which and how many genes a chromosome contains, and the kind of protein that they produce. However, the Genome map doesn’t give information about which of these genes are turned on or off in different tissues, in various stages of development or in pathological situations. This kind of information is obtained by expression maps: thanks to these maps we have the knowledge of which genes are important for the formation of a particular tissue or organ. Furthermore, they help us to identify the genes that are expressed in a different way between a healthy tissue and a pathological tissue. While the genomic maps analyse the DNA, the expression studies analyse the RNA, a molecule similar to DNA, produced only by active genes. Each active gene produces a different type of RNA: the analysis of the presence of each type of RNA tells us if the corresponding gene is “on” (= expressed) or “off” (= not expressed). Furthermore, the amount of RNA indicates the expression degree of the corresponding gene.
Is
it possible to have a practical example of these experiments?
The following images, taken from the study published in Nature, show some examples of the expression atlas.
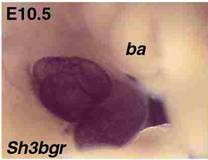 Fig 1.
The dark purple mass in the middle of the image (carried out on a low power
microscope) is a rodent heart, during embryonic development, in which a
chromosome 21 gene, named Sh3bgr, has been “labelled”. The dark purple signal
indicates that the gene is strongly expressed in the heart, but is “off” in the
nearby tissues. From this experiment we deduce that the gene in question might
have a role in heart development.
Fig 1.
The dark purple mass in the middle of the image (carried out on a low power
microscope) is a rodent heart, during embryonic development, in which a
chromosome 21 gene, named Sh3bgr, has been “labelled”. The dark purple signal
indicates that the gene is strongly expressed in the heart, but is “off” in the
nearby tissues. From this experiment we deduce that the gene in question might
have a role in heart development.
aorta atria ventricles
![]()
![]()
![]()
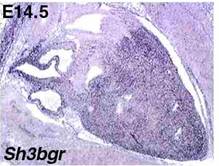 Fig 2.
In this picture, the same heart (the darker mass occupying almost the whole
image) is seen at higher power. In this way it is possible to observe that the
gene is expressed in both the ventricles and the atria but not in the aorta.
Fig 2.
In this picture, the same heart (the darker mass occupying almost the whole
image) is seen at higher power. In this way it is possible to observe that the
gene is expressed in both the ventricles and the atria but not in the aorta.
What
is Down Syndrome?
Down Syndrome is the most frequent genetic cause of mental retardation, with a frequency of 1 child affected out of 700. It is also known as trisomy 21, as it is caused by the presence of 3 copies of chromosome 21, instead of 2. The symptoms of Down Syndrome are complex and involve various organs: the most frequent characteristics are mental retardation, low stature, short limbs, oblique palpebral fissures, protruding tongue, cardiac defects (present in 40% of the patients), a higher incidence of leukemia and defects of the immune system. At present, no treatment to cure or prevent the syndrome does exist. The current treatments, though important, can only monitor and correct some of the symptoms, such as cardiac defects. Rehabilitation therapy, instead, aims to obtain a harmonic development and a good scholastic, social and professional integration.
What
is the importance of this study for the research on Down syndrome and other
genetic diseases linked to chromosome 21?
Down syndrome is caused by the presence of an extra copy of chromosome 21. In other words, people affected with the syndrome have a supernumerary copy of all the genes contained in this chromosome. Still, nobody knows exactly which and how many of these genes are important to determine the symptoms and signs of the syndrome. There is no evidence that these genes are responsible for one or more symptoms of this disease, or even for mental retardation.
This expression atlas is an important instrument for connecting the expression of these genes with the various symptoms of Down syndrome. It also gives us insight in which and how many genes are involved in the origins of this disease.
For example, the study has revealed that some of the genes of chromosome 21 are “on” in particular regions of the heart during embryonic development: therefore these genes may have a role in congenital heart defects found in Down patients. In the same way, some genes, which are “on” in the embryonic limb buds, might be involved in hand and feet anomalies typical of the syndrome. Finally, a set of genes of chromosome 21 is expressed in the brain, suggesting that they may play a role in mental retardation.
As already said, chromosome 21 contains at least 30 genes involved in hereditary diseases other than Down syndrome. The expression atlas will give us a more precise insight on the role of these genes, which, to a large extent, is still unknown.
Does
the study allow practical application for treatment or diagnosis of Down Syndrome or other diseases?
It is necessary to explain that this study represents a basic research for which no immediate application has been foreseen in the diagnosis or treatment of Down syndrome or other diseases. The study published in the prestigious scientific journal Nature (http://www.nature.com/nature) will surely help in understanding which genes of chromosome 21 have a key-role in the origins of the various symptoms characterizing Down Syndrome. The identification of these genes does not mean that a therapy will be immediately found for the disease, unfortunately, and a definite solution is still far away. Nevertheless, the study represents a major advancement for research on Down syndrome and on the other diseases linked to chromosome 21. For many years we were left in the dark; today, thanks to the results of the Genome project and the expression atlas, researchers have a guide in hand to better understand the genetic basis of Down syndrome and of the other diseases linked to chromosome 21.